| Issue |
Wuhan Univ. J. Nat. Sci.
Volume 31, Number 1, February 2026
|
|
|---|---|---|
| Page(s) | 1 - 9 | |
| DOI | https://doi.org/10.1051/wujns/2026311001 | |
| Published online | 06 March 2026 | |
Deep Learning and Intelligent Perception
CLC number: TP391.41
MS-RWKV-UNet: Multi-Head Scan Receptance Weighted Key Value UNet for Medical Image Segmentation
基于多头扫描策略加权键值(RWKV)网络的医学图像分割方法
1
School of Computer Science and Technology, Hangzhou Dianzi University, Hangzhou 310018, Zhejiang, China
(杭州电子科技大学 计算机学院,浙江 杭州 310018)
2
School of Computer Science and Cyber Engineering, Guangzhou University, Guangzhou 510014, Guangdong, China
(广州大学 计算机科学与网络工程学院,广东 广州 510006)
† Corresponding author. E-mail: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
6
September
2025
Abstract
The Transformer has achieved great success in the field of medical image segmentation, but its quadratic computational complexity limits its application in dense medical image prediction. Recently, the receptance weighted key value (RWKV) architecture has garnered widespread attention due to its linear computational complexity and its capability of parallel computation during training. Despite the RWKV model's proficiency in addressing long-range modeling tasks with linear computational complexity, most current RWKV-based approaches employ static scanning patterns. These patterns may inadvertently incorporate biased prior knowledge into the model's predictions. To address this challenge, we propose a multi-head scan strategy combined with padding methods to effectively simulate spatial continuity in 2D images. Within the Feature Aggregation Attention (FAA) module, asymmetric convolutions are designed to aggregate 1D sequence features along a single dimension, thereby expanding effective receptive fields while preserving structural sparsity. Additionally, panoramic token shift (P-Shift) effectively models local dependency relationships by moving tokens from a wide receptive field. Extensive experiments conducted on the ISIC17/18 and ACDC datasets demonstrate that our method exhibits superior performance in dense medical image prediction tasks.
摘要
尽管Transformer架构在医学图像分割领域取得了显著成果,但其自注意力机制固有的二次计算复杂度限制了在密集预测任务中的应用。近年来,RWKV架构因其线性计算复杂度及训练时的高并行能力受到广泛关注。尽管RWKV模型能够以线性计算复杂度有效处理远程建模任务,但当前基于RWKV的方法多依赖静态扫描模式,容易引入有偏的先验知识,影响模型泛化性能。为应对这一挑战,我们提出结合填充方法的多头扫描策略,以更好地模拟二维图像中的空间连续性。在特征聚合注意力(FAA)模块中,通过设计异构卷积沿单一维度融合一维序列特征,在保持结构稀疏性的同时扩展有效感受野。此外,P-Shift通过宽感受野内的token移动增强局部依赖建模。在ISIC和ACDC数据集上的大量实验表明,所提出方法在多项密集预测任务中均优于现有基线模型,展现出更高的分割精度和鲁棒性。
Key words: multi-head scan receptance weighted key value (RWKV) / asymmetric convolution / panoramic token shift (P-Shift) / medical image segmentation
关键字 : 多头扫描RWKV / 异构卷积 / P-Shift / 医学图像分割
Cite this article:JIANG Dong, JI Zhongping, FANG Meie. MS-RWKV-UNet: Multi-Head Scan Receptance Weighted Key Value UNet for Medical Image Segmentation[J]. Wuhan Univ J of Nat Sci, 2026, 31(1): 1-9.
Biography: JIANG Dong, male, Master candidate, research direction: computer vision and medical image segmentation, etc. E-mail: This email address is being protected from spambots. You need JavaScript enabled to view it.
Foundation item: Supported by Zhejiang Provincial Natural Science Foundation of China (LY22F020025) and the National Natural Science Foundation of China( 62072126)
© Wuhan University 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
0 Introduction
Precise and efficient medical image segmentation is paramount in medical image analysis [1-2]. However, conventional manual segmentation is labor-intensive, time-consuming, and prone to inter-expert variability [3-4]. This necessitates automated approaches, particularly deep learning algorithms, to accurately delineate organs or pathological regions. Such methods are crucial for facilitating accurate, rapid, and consistent diagnoses for clinicians and researchers[5]. In recent years, the proliferation of advanced deep learning architectures, including Convolutional Neural Networks (CNNs), Vision Transformers (ViTs), and Mamba, has led to substantial breakthroughs in medical image segmentation. Nonetheless, despite their notable benefits, each architecture faces inherent performance constraints dictated by its intrinsic design attributes [6]. CNNs, relying on local convolutional kernels, inherently struggle to capture long-range dependencies, which can lead to suboptimal feature extraction and segmentation outcomes. Conversely, ViTs, while adept at global modeling, are hampered by quadratic computational complexity, limiting their efficiency in dense prediction tasks. Furthermore, Mamba-based models encounter challenges in achieving an optimal effective receptive field [7] when converting 2D image data into 1D sequential formats.
The Receptance Weighted Key Value (RWKV) model[8] introduces the WKV attention mechanism alongside token shift layers, facilitating linear computational complexity within global attention mechanisms while effectively capturing local dependencies. Despite the innovative efforts to extend the application of RWKV to visual domains, challenges persist in the direct adaptation of RWKV for dense image prediction tasks, including the nuanced field of medical image segmentation. This primarily stems from the inherent incompatibility between the causal sequential modeling capabilities of the RWKV architecture and the 2D spatial structure of images. Tailored for 1D sequences modeling, RWKV is not inherently suited for direct application to model 2D image tokens. Prior research has addressed this issue by flattening 2D tokens into 1D sequences via a computational hierarchy[9], such as row-column-major ordering, where each row's end is immediately followed by the next row's start. However, this approach ignores the preservation of spatial continuity[10], thereby compromising the integrity of the intrinsic structural information of the image.
This paper introduces an optimized medical image segmentation framework, termed Multi-head Scan RWKV (MS-RWKV), which is an adaptation of the RWKV architecture [8] for 2D visual tasks. The proposed model preserves the fundamental architecture and inherent benefits of RWKV, while incorporating essential modifications tailored for the segmentation of 2D medical images. 1) Our multi-head scanning module reduces the unidirectional causal bias among image patches, enabling more balanced global receptive field computation. We strategically insert padding tokens between scan-sequence elements that are spatially disconnected, preserving 2D structural continuity. 2) Building on the GhostNetV2 [11] architecture, we design a feature aggregation block that captures local spatial context while strengthening correlations within 1D sequences generated along specific scan directions, using asymmetric convolutions. 3) This novel mechanism broadens token semantics by aggregating multi-scale features from wide receptive fields, effectively addressing orientation sensitivity in 2D images. During training, panoramic token shift (P-Shift) employs structural reparameterization to learn adaptive token shifting across diverse contexts.
We rigorously evaluated our proposed MS-RWKV model through comprehensive experiments on skin lesion segmentation and multi-organ segmentation tasks, showcasing its superior performance and high efficiency in the field of medical image segmentation. Ad-ditionally, we conducted ablation studies to validate the performance of our design. An extensive array of experimental outcomes underscores the robust potential of our model for image segmentation applications.
The main contributions of this study are as follows:
• We introduced the MS-RWKV framework, adapting the RWKV model for application in medical image segmentation tasks. This adaptation has demonstrated promise as an enhanced solution for more precise and effective image segmentation.
• We integrated a novel multi-head scan mechanism, augmented with padding strategies, into the RWKV architecture. This innovation effectively bridges the divide between 1D sequences processing and 2D image traversal.
• During the conversion of 2D images to 1D sequences, we integrate the Feature Aggregation Attention (FAA) module. The asymmetric convolution within this module extracts features that are particularly advantageous for subsequent processing of 1D sequences.
1 Methodology
The RWKV model[12], drawing from natural language processing, combines the parallel training efficiency of transformers[13] with the sequential inference capabilities of Recurrent Neural Networks (RNNs)[14]. The architecture of the RWKV model comprises a series of stacked blocks, each of which integrates time-mixing and channel-mixing blocks, both of which feature recurrent structures.
Time-Mixing Block. This block is engineered to augment the modeling capacity for dependencies and patterns within sequential data. Given an input sequence 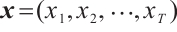 , where T represents the length of the input features after convolutional subsampling, the output sequence
, where T represents the length of the input features after convolutional subsampling, the output sequence 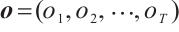 of the time-mixing block is computed as follows:
of the time-mixing block is computed as follows:
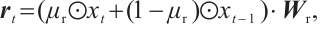 (1)
(1)
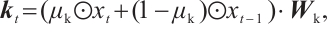 (2)
(2)
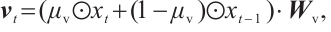 (3)
(3)
 (4)
(4)
where 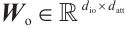 is the output projection matrix,
is the output projection matrix,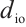 is the input/output size, and
is the input/output size, and 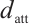 is the RWKV time-mixing block size.
is the RWKV time-mixing block size. 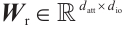 ,
, 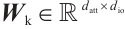 and
and 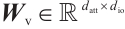 are the projection matrices for the receptance, key, and value, respectively.
are the projection matrices for the receptance, key, and value, respectively. 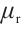 ,
, 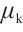 and
and 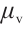 are time-mixing factors for the receptance, key, and value, respectively. The values of
are time-mixing factors for the receptance, key, and value, respectively. The values of 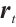 ,
, 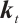 , and
, and 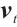 are calculated through linear interpolation between the current input and the input from the previous time step. This block applies a non-linear activation function σ to the receptance vector
are calculated through linear interpolation between the current input and the input from the previous time step. This block applies a non-linear activation function σ to the receptance vector 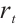 , and then combines the resulting values with the hidden state
, and then combines the resulting values with the hidden state 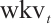 through element-wise multiplication.
through element-wise multiplication.
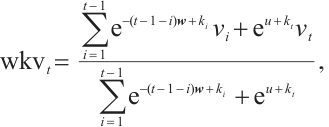 (5)
(5)
where 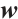 is the channel-wise time decay vector for the previous input,
is the channel-wise time decay vector for the previous input, 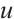 is the special weighting factor applied to the current input, and
is the special weighting factor applied to the current input, and 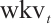 is the weighted summation of the input in the interval [1, t]. The hidden states Eq. (5) can be computed recursively as follows:
is the weighted summation of the input in the interval [1, t]. The hidden states Eq. (5) can be computed recursively as follows:
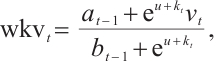 (6)
(6)
where 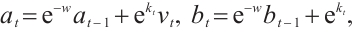 and
and 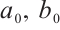 are zero-initialized.
are zero-initialized.
Channel-Mixing Block. This block is specifically engineered to enhance the feature representations propagated from the time-mixing block through a series of robust non-linear transformations. Given the input sequence 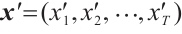 , the specific process of the block is:
, the specific process of the block is:
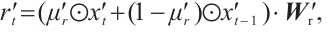 (7)
(7)
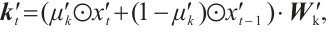 (8)
(8)
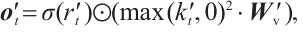 (9)
(9)
where 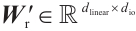 and
and 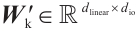 are the projection matrices for the receptance and key, respectively.
are the projection matrices for the receptance and key, respectively. 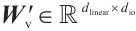 is the channel-mixing matrix, and
is the channel-mixing matrix, and 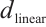 is the RWKV time-mixing block size.
is the RWKV time-mixing block size. 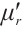 and
and 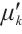 are time-mixing factors for the receptance and key, respectively. The channel-mixing block operates causally, as the computation of ot′ is contingent solely on xt′ and xt-1′ . Intuitively, this amplification process enhances the representations of historical information.
are time-mixing factors for the receptance and key, respectively. The channel-mixing block operates causally, as the computation of ot′ is contingent solely on xt′ and xt-1′ . Intuitively, this amplification process enhances the representations of historical information.
2 Architecture of MS-RWKV
The RWKV[15, 12] architecture, originally conceived for processing 1D sequences, encoun-ters some limitations when tasked with learning from 2D data structures. To address these challenges, we introduce novel modules that enhance the RWKV's capability to effectively process 2D image data.
Overall architecture. The architecture of the MS-RWKV is depicted in Fig. 1(a). The MS-RWKV incorporates a four-stage hierarchical backbone with skip connections. Given an input image I, we first partition the feature map 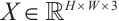 into 2D patches via the non-overlapping patch embedding layer, with the channel dimension projected into c dimensions. As illustrated in Fig.1 (b), each stage's MS-RWKV module is tasked with subsequently extracting feature representations at varying levels from the input image. Within these modules, four distinct multi-head scanning trajectories are utilized to linearize the patch tokens into sequences X= [S1, S2, S3, S4], where Sn is the sequence after the n-th scan path. The MS-RWKV module achieves competitive performance by computing global and local attention with linear complexity for input sequences. Following the decoding phase, the image resolution is restored to its original size through the final projection layer, enabling pixel-accurate segmentation. A comprehensive exposition of our architectural design principles and methodological implementations will be systematically delineated in the following sections.
into 2D patches via the non-overlapping patch embedding layer, with the channel dimension projected into c dimensions. As illustrated in Fig.1 (b), each stage's MS-RWKV module is tasked with subsequently extracting feature representations at varying levels from the input image. Within these modules, four distinct multi-head scanning trajectories are utilized to linearize the patch tokens into sequences X= [S1, S2, S3, S4], where Sn is the sequence after the n-th scan path. The MS-RWKV module achieves competitive performance by computing global and local attention with linear complexity for input sequences. Following the decoding phase, the image resolution is restored to its original size through the final projection layer, enabling pixel-accurate segmentation. A comprehensive exposition of our architectural design principles and methodological implementations will be systematically delineated in the following sections.
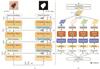 |
Fig. 1 (a) The overall architecture of MS-RWKV. (b) The core of MS-RWKV block |
Feature Aggregation Module. The RWKV[12] model, initially tailored for 1D input sequences, encounters challenges in preserving local dependency relationships when applied to 2D image data, which impedes its ability to capture local fine-grained details[16]. Building upon GhostNetV2's[11] approach to capturing local details, we develop a novel feature aggregation module, Feature Aggregation Attention (FAA), to enhance feature aggregation in dense prediction tasks. This module is engineered to enhance local feature extraction capability by expanding effective receptive fields while preserving parameter sparsity through grouped convolutions. FAA consists of two Multi-scale Convolutional Modules (MCM) and an Activation Module (AM). It can be described as follows:
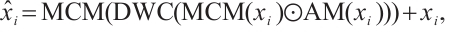 (10)
(10)
where xi and 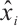 represent the input and output tensors of the module in the i-th stage, DWC is a depth-wise convolution with a kernel size of 3 × 3.
represent the input and output tensors of the module in the i-th stage, DWC is a depth-wise convolution with a kernel size of 3 × 3.
As illustrated in Fig. 2 , the input x is partitioned into four subcomponents x = (X1, X2, X3, X4), which are processed through parallel convolutional pathways with asymmetric kernel dimensions. The resultant features are concatenated to form the final output, achieving multi-granularity feature fusion that captures both local details and global context. Crucially, the asymmetric convolutions (kernel size: 1 × KH , KW × 1) explicitly model directional spatial correlations, thereby significantly facilitating sequential scanning in the subsequent RWKV module. The AM branch implements a module consisting of two linear layers and an activation function. FAA leverages the features expanded by the first MCM module, which are then modulated by the AM branch, augmenting the model's expressive power. The enhanced features are subsequently fed into the second MCM module to restore the original feature information for output, effectively aggregating surrounding information. This method effectively mitigates the inherent limitations of the flattening approach.
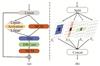 |
Fig. 2 (a) The diagrams of blocks in feature aggregation module. (b) Multi-scale convolutional module |
Scan patterns. The multi-head scan mechanism, which involves the parallel extraction and integration of features across various divergent scanning trajectories [17-18], enables the capture of a global receptive field and the modeling of long-range dependencies. This module also draws inspiration from the research on multi-head scan presented in UltraLight VM-UNet[19] and MHS-VM[20], and we conducted correlation experiments to explore the implications and applications of these methods. The details of the aforementioned process can be formulated as follows:
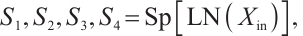 (11)
(11)
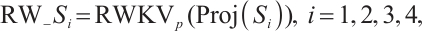 (12)
(12)
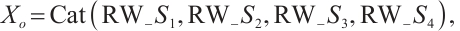 (13)
(13)
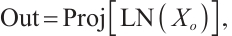 (14)
(14)
where LN is the LayerNorm, Sp is the Split operation, RWKVp is the RWKV operation with padding, Cat is the concatenation operation, and Proj is the Projection operation. The information gathered from the four branches is then merged and cycled through the subsequent model module. Within each layer, consecutive modules integrate various scanning approaches, thereby enhancing the model's generalization capabilities[21]. Ultimately, we opted for a parallel 4-head scan approach to uniformly decompose the depth feature Xin into four sequences [S1, S2, S3, S4]. In Eqs.(12) and (13), each branch RWKVp with padding p independently extracts pertinent information, which is subsequently aggregated and output through a concatenation strategy. To address the spatial discontinuity resulting from the flattening of the image into 1D sequences, we insert padding tokens at the end of each row and column to detect the end-of-line signal. We execute both row-major and column-major scans in both forward and reverse directions, as delineated in Fig.3. To preserve the resolution of the original image, we remove padding tokens prior to concatenation, thereby restoring the image to its input size. Finally, the four features are concatenated to form the composite feature Xo, which is subsequently processed by LayerNorm and Projection operations to yield the Out.
 |
Fig. 3 Illustrations of the scan head block Note: After padding the image, one of the four scanning methods is selected to flatten the image into 1D sequences. |
Token shift. The token shift mechanism in RWKV [8] was initially introduced to mitigate the misalignment between the 1D decay of attention and the 2D adjacency in images. However, current token shift methods, including the unidirectional token shift (Uni-shift) in RWKV [8] and the quadridirectional token shift (Quad-shift) in Vision-RWKV[22], collect information from limited directions and fail to account for the comprehensive spatial continuity inherent in 2D imagery. To address this, we employ a panoramic token shift (P-Shift) mechanism, integrating it into time-mixing and channel-mixing blocks in the RWKV module. In our approach, we harness a suite of deep convolutional layers equipped with convolutional kernels of diverse sizes to facilitate the extraction and fusion of features. This strategy effectively aggregates information from multiple spatial orientations, enhancing the model's capacity to capture richer spatial features[23]. The mechanism of P-Shift can be formulated as follows:
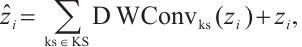 (15)
(15)
where 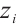 and
and 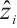 represent the input and output tensors of the module in the i-th tage. DWConvks denotes a depthwise convolution with a kernel size of ks. KS defines a set of parallel convolution kernels with values of {1 × 1, 3 × 3, 5 × 5}. This mechanism utilizes a multi-branch architecture during training to capture local contextual information with an expanded visual receptive field. For testing, the multi-branch structure [24] is consolidated into a single branch using a 5×5 convolution kernel, thereby enhancing the model's inference efficiency and reducing parameter count.
represent the input and output tensors of the module in the i-th tage. DWConvks denotes a depthwise convolution with a kernel size of ks. KS defines a set of parallel convolution kernels with values of {1 × 1, 3 × 3, 5 × 5}. This mechanism utilizes a multi-branch architecture during training to capture local contextual information with an expanded visual receptive field. For testing, the multi-branch structure [24] is consolidated into a single branch using a 5×5 convolution kernel, thereby enhancing the model's inference efficiency and reducing parameter count.
3 Experiments and Results
3.1 Datasets and Parameter
We conducted an extensive performance evaluation of our framework across three distinct open-source medical image segmentation datasets: ISIC17[25], ISIC18 [26] and ACDC[27]. These benchmark datasets are widely recognized for their pivotal role in advancing medical image segmentation research. In our experimental setup, we implemented the MS-RWKV model using PyTorch 2.0 and performed all training on an NVIDIA GeForce RTX 3090Ti GPU. To enhance the robustness of our model, we incorporated data augmentation techniques, including random flipping and rotation. The training was conducted with a batch size of 32 over 300 epochs. We opted for the AdamW optimizer for network training, complemented by a cosine annealing schedule for learning rate decay. Given the heterogeneity in difficulty across different datasets, we fine-tuned the hyperparameters accordingly. For the ACDC dataset, we initialized the learning rate at 5 × 10-4 and set the weight decay to 1 × 10-4. For the remaining datasets, we standardized the learning rates at 1 × 10-3 and weight decays at 1 × 10-5.To ensure a rigorous and comprehensive performance evaluation, we adopted a standardized set of quantitative metrics, including the Mean Intersection over Union (mIoU), the Dice Similarity Coefficient (DSC), and Accuracy (Acc).
3.2 Main Results
To verify the effectiveness of our proposed method, we performed a comparative analysis of the MS-RWKV model against several state-of-the-art models.
Table 1 presents our method's performance compared with various approaches on ISIC17 and ISIC18 datasets, where our proposed MS-RWKV achieved the best average mIoU of 83.27% and 81.52%. Specifically, compared with Mamba-based methods (such as H-vmunet[28]), our method improved the mIoU by 1.22% and 0.92%, respectively, and by 0.56% and 0.47% compared with RWKV-based methods (such as RWKV-UNet[29]). As shown in Table 2, compared with other methods, our proposed method achieved the best average DSC of 91.85% on the ACDC dataset.
The quantitative results presented in the tables demonstrate that our method achieves superior performance compared to state-of-the-art approaches for 2D medical image segmentation tasks.
Comparative experimental results on the ISIC17 and ISIC18 dataset (unit:%)
Performance comparison with state-of-the-art methods on the ACDC dataset for right ventricle (RV), Myocardium (Myo), and left ventricle (LV) segmentation (unit:%)
3.3 Ablation Studies
We conducted comprehensive ablation studies on the ISIC18 dataset to validate the effectiveness of the multi-head scan, feature aggregation attention, and P-shift components. The results are shown below.
Multi-Head Scan with Padding. In our ablation studies, we adopt four parallel scan heads for hierarchical feature extraction from 2D image data. To systematically evaluate the impact of architectural components, we conducted comparative experiments with two distinct configurations: 1) four scan heads without padding mechanisms, and 2) three scan heads with strategic padding implementation. The quantitative results, as detailed in Table 3, demonstrate that increasing the number of scan heads leads to enhanced model capacity and improved feature representation.
Token Shift. To explore the effectiveness of the proposed P-Shift, we compared its performance with Uni-Shift in RWKV[8] and Quad-Shift in Vision-RWKV[22]. The ablation studies presented in Table 4 demonstrate that our proposed P-Shift mechanism coupled with reparameterization significantly enhances the local feature extraction capability of the token shift operation. Our novel token shift architecture effectively exploits the inherent spatial correlations within 2D visual feature maps, facilitating multi-directional feature propagation and adaptive feature aggregation across diverse spatial orientations.
Feature Aggregation Attention. To systematically investigate the impact of various architectural components preceding the MS-RWKV Block, we conducted a series of ablation experiments. Our assessment focused on comparing the performance of two distinct baseline modules (FFN and GhostNetV2) against our proposed FAA module. As presented in Table 5, the proposed FAA module achieves superior performance, surpassing FFN by 1.32% and GhostNetV2 by 0.54% in mIoU.
We conducted comprehensive ablation studies on ISIC18 to evaluate individual components. Table 6 summarizes ablation results, where the last row shows the full model's performance and the preceding rows quantify the impact of module removal. A comparative analysis between the full model and the configurations lacking the FAA reveals that FAA contributes most significantly to the overall performance.
Ablation study on number of scan heads and padding
Ablation study on P-Shift (unit:%)
Ablation study on FAA (unit:%)
Ablation study on P-shift and FAA (unit:%)
3.4 Visualization
Qualitative analysis of ISIC 2018 results (Fig. 4) demonstrates that our MS-RWKV architecture facilitates the feature integration of semantic information and finer-grained features. Quantitative experiments confirm that our proposed framework achieves significant gains over the VM-UNet baseline, particularly in capturing subtle texture variations as confirmed by quantitative metrics. These visualizations further validate that MS-RWKV, as an RWKV-based model, holds significant potential in the field of medical image segmentation.
 |
Fig. 4 The visual comparison of segmentation results of ours model and other segmentation methods against ground truth on ISIC2018 dataset |
4 Conclusion
In this paper, we propose MS-RWKV, which extends the foundational architecture of the RWKV model. MS-RWKV enhances RWKV through three innovations: multi-head scanning with adaptive padding for 2D coherence, P-Shift for spatial dependencies, and feature aggregation attention for multi-scale fusion. Comprehensive empirical evaluations across diverse medical image segmentation benchmarks demonstrate that our MS-RWKV architecture achieves superior performance compared to state-of-the-art approaches. We further plan to extend MS-RWKV to tasks beyond segmentation, such as cross-modal registration and high-fidelity reconstruction.
References
- Bai W J, Suzuki H, Huang J, et al. A population-based phenome-wide association study of cardiac and aortic structure and function[J]. Nature Medicine, 2020, 26(10): 1654-1662. [Google Scholar]
- Fatma K, Benaissa I, Zitouni A, et al. Assessing the performance of U-Net in 3D medical image segmentation[C]//2024 8th International Conference on Image and Signal Processing and Their Applications (ISPA). New York: IEEE, 2024: 1-6. [Google Scholar]
- Jungo A, Meier R, Ermis E, et al. On the effect of inter-observer variability for a reliable estimation of uncertainty of medical image segmentation[C]//Medical Image Computing and Computer Assisted Intervention – MICCAI 2018. Cham: Springer-Verlag, 2018: 682-690. [Google Scholar]
- Joskowicz L, Cohen D, Caplan N, et al. Inter-observer variability of manual contour delineation of structures in CT[J]. European Radiology, 2019, 29(3): 1391-1399. [Google Scholar]
- Tang H, Chen X M, Liu Y, et al. Clinically applicable deep learning framework for organs at risk delineation in CT images[J]. Nature Machine Intelligence, 2019, 1(10): 480-491. [Google Scholar]
- Chen J N, Mei J R, Li X H, et al. TransUNet: Rethinking the U-Net architecture design for medical image segmentation through the lens of transformers[J]. Medical Image Analysis, 2024, 97: 103280. [Google Scholar]
- Yang Z W, Li J Y, Zhang H, et al. Restore-RWKV: Efficient and effective medical image restoration with RWKV[J]. IEEE Journal of Biomedical and Health Informatics, 2025, 28(3): 1484-1493. [Google Scholar]
- Peng B, Alcaide E, Anthony Q, et al. RWKV: Reinventing RNNs for the transformer era[C]//Findings of the Association for Computational Linguistics: EMNLP 2023. Stroudsburg: ACL, 2023: 14048-14077. [Google Scholar]
- Tsai T Y, Lin L, Hu S, et al. UU-mamba: Uncertainty-aware U-mamba for cardiac image segmentation[C]//2024 IEEE 7th International Conference on Multimedia Information Processing and Retrieval (MIPR). New York: IEEE, 2024: 267-273. [Google Scholar]
- Duan X J, Shi M C, Wang J M, et al. Segmentation of the aortic dissection from CT images based on spatial continuity prior model[C]//2016 8th International Conference on Information Technology in Medicine and Education (ITME). New York: IEEE, 2016: 275-280. [Google Scholar]
- Tang Y H, Han K, Guo J Y, et al. GhostNetV2: Enhance cheap operation with long-range attention[C]//Advances in Neural Information Processing Systems 35 (NeurIPS 2022). 2022: 1-12. [Google Scholar]
- Li Z Y, Xia T Y, Chang Y, et al. A survey of RWKV[EB/OL]. [2024-01-12]. arXiv:2412.14847. [Google Scholar]
- Dosovitskiy A, Beyer L, Kolesnikov A, et al. An image is worth 16x16 words: Transformers for image recognition at scale[C]//International Conference on Learning Representations (ICLR), 2021: 1-22. [Google Scholar]
- Graves A. Long short-term memory[M]//Supervised Sequence Labelling with Recurrent Neural Networks. Berlin: Springer-Verlag, 2012: 37-45. [Google Scholar]
- Zhou L, Xiao Z L, Ning Z P. RWKV-based encoder-decoder model for code completion[C]//2023 3rd International Conference on Electronic Information Engineering and Computer (EIECT). New York: IEEE, 2023: 425-428. [Google Scholar]
- Huang T, Pei X H, You S, et al. LocalMamba: Visual state space model with windowed selective scan[C]//Computer Vision–ECCV 2024 Workshops. LNCS15633. Cham: Springer-Verlag, 2025: 13-32. [Google Scholar]
- Cai Z F, Fan Y L, Zhu M W, et al. Ultra-lightweight network for medical image segmentation inspired by bio-visual interaction[J]. IEEE Transactions on Circuits and Systems for Video Technology, 2025, 35(4): 3486-3497. [Google Scholar]
- Peng B, Chen K, Xu Y, et al. RSMamba: Remote sensing image classification with state space model[J]. IEEE Geoscience and Remote Sensing Letters, 2024, 21: 1-5. [Google Scholar]
- Wu R K, Liu Y H, Liang P C, et al. UltraLight VM-UNet: Parallel vision mamba significantly reduces parameters for skin lesion segmentation[J]. Patterns, 2025, 6(7): 101298. [Google Scholar]
- Ji Z P. MHS-VM: Multi-head scanning in parallel subspaces for vision Mamba[EB/OL]. [2024-01-12]. arXiv:2406.05992. [Google Scholar]
- Chen S Q, Zhong X, Dorn S, et al. Improving generalization capability of multiorgan segmentation models using dual-energy CT[J]. IEEE Transactions on Radiation and Plasma Medical Sciences, 2022, 6(1): 79-86. [Google Scholar]
- Duan Y C, Wang W Y, Chen Z, et al. Vision-RWKV: Efficient and scalable visual perception with RWKV-like architectures[C]//Proceedings of the International Conference on Learning Representations (ICLR), 2025: 1-23. [Google Scholar]
- Kaleybar J M, Saadat H, Khaloo H. Capturing local and global features in medical images by using ensemble CNN-Transformer[C]//Proceedings of the 13th International Conference on Computer and Knowledge Engineering (ICCKE). New York: IEEE, 2023: 1-6. [Google Scholar]
- Liu Y S, Zhao Y J, Wang M H, et al. MBD-net: Multi-branch dilated convolutional network with cyst discriminator for renal multi-structure segmentation[C]//45th Annual International Conference of the IEEE Engineering in Medicine & Biology Society. New York: IEEE, 2023: 1-4. [Google Scholar]
- Codella N C F, Gutman D, Celebi M E, et al. Skin lesion analysis toward melanoma detection: A challenge at the 2017 International symposium on biomedical imaging (ISBI)[C]//IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018). New York: IEEE, 2018: 168-172. [Google Scholar]
- Codella N, Rotemberg V, Tschandl P, et al. Skin lesion analysis toward melanoma detection 2018: A challenge hosted by the international skin imaging collaboration (ISIC)[C]//Proceedings of the Medical Image Computing and Computer Assisted Intervention . 2019: 168-172. [Google Scholar]
- Bernard O, Lalande A, Zotti C, et al. Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: Is the problem solved?[J]. IEEE Transactions on Medical Imaging, 2018, 37(11): 2514-2525. [Google Scholar]
- Wu R K, Liu Y H, Liang P C, et al. H-VMUnet: High-order vision mamba unet for medical image segmentation[J]. Neurocomputing, 2025:129447. [Google Scholar]
- Jiang J T, Zhang J N, Liu W X, et al. RWKV-UNet: Improving UNet with Long-Range Cooperation for Effective Medical Image Segmentation[EB/OL]. [2024-01-12]. arXiv:2501.08458. [Google Scholar]
- Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation[C]//Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. LNCS9351. Cham: Springer-Verlag, 2015: 234-241. [Google Scholar]
- Ruan J C, Xiang S H, Xie M Q, et al. MALUNet: A multi-attention and light-weight UNet for skin lesion segmentation[C]//2022 IEEE International Conference on Bioinformatics and Biomedicine. New York: IEEE, 2022: 1150-1156. [Google Scholar]
- Zhang Y D, Liu H Y, Hu Q. TransFuse: Fusing transformers and CNNs for medicalimage segmentation[C]//Proceedings of the 24th International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham: Springer-Verlag, 2021: 1150-1156. [Google Scholar]
- Ruan J C, Li J C, Xiang S C. VM-UNet: Vision Mamba UNet for Medical Image Segmentation[EB/OL]. [2024-01-12]. arXiv:2402.02491. [Google Scholar]
- Oktay O, Schlemper J, Le Folgoc L, et al. Attention U-Net: Learning where to look for the pancreas[J]. Proceedings of the Medical Imaging with Deep Learning (MIDL), 2019, 53: 197-207. [Google Scholar]
- Cao H, Wang Y Y, Chen J, et al. Swin-Unet: Unet-like pure transformer forMedical image segmentation[C]//Computer Vision – ECCV 2022 Workshops. Cham: Springer-Verlag, 2023: 205-218. [Google Scholar]
- Huang X H, Deng Z F, Li D D, et al. MISSFormer: An effective transformer for 2D medical image segmentation[J]. IEEE Transactions on Medical Imaging, 2023, 42(5): 1484-1494. [Google Scholar]
- Hatamizadeh A, Tang Y C, Nath V, et al. UNETR: Transformers for 3D medical image segmentation[C]//2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). New York: IEEE, 2022: 1748-1758. [Google Scholar]
All Tables
Performance comparison with state-of-the-art methods on the ACDC dataset for right ventricle (RV), Myocardium (Myo), and left ventricle (LV) segmentation (unit:%)
All Figures
 |
Fig. 1 (a) The overall architecture of MS-RWKV. (b) The core of MS-RWKV block |
| In the text | |
 |
Fig. 2 (a) The diagrams of blocks in feature aggregation module. (b) Multi-scale convolutional module |
| In the text | |
 |
Fig. 3 Illustrations of the scan head block Note: After padding the image, one of the four scanning methods is selected to flatten the image into 1D sequences. |
| In the text | |
 |
Fig. 4 The visual comparison of segmentation results of ours model and other segmentation methods against ground truth on ISIC2018 dataset |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


