| Issue |
Wuhan Univ. J. Nat. Sci.
Volume 31, Number 2, April 2026
|
|
|---|---|---|
| Page(s) | 146 - 156 | |
| DOI | https://doi.org/10.1051/wujns/2026312146 | |
| Published online | 13 May 2026 | |
Aquatic Ecology and Water Environment Safety
CLC number: X524
Antibiotic Resistance Genes in Aquatic Environments: Sources, Transmission, and Influencing Factors from a One Health Perspective
水生环境中抗生素抗性基因:基于“One Health”视角的来源、传播及影响因素
1
Public Health College, Mudanjiang Medical University, Mudanjiang 157011, Heilongjiang, China
(牡丹江医科大学 公共卫生学院, 黑龙江 牡丹江 157011)
2
Heilongjiang River Basin Fishery Ecological Environment Monitoring Center, Ministry of Agriculture and Rural Affairs, Heilongjiang River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Harbin 150010, Heilongjiang, China
(中国水产科学研究院 黑龙江水产研究所 农业农村部黑龙江流域渔业生态环境监测中心, 黑龙江 哈尔滨 150010)
3
Fisheries Research Institute, Sichuan Academy of Agricultural Sciences (Sichuan Fisheries Research Institute), Chengdu 611731, Sichuan, China
(四川省农业科学院 水产研究所(四川省水产科学研究院), 四川 成都 611731)
4
Fisheries College, Sichuan Agricultural University, Chengdu 611130, Sichuan, China
(四川农业大学 水产学院, 四川 成都 611130)
† Corresponding author. E-mail: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
30
October
2025
Abstract
Antibiotic resistance genes (ARGs) are emerging environmental contaminants that pose significant threats to public health due to their persistence, migration and dissemination in aquatic environments. This review systematically summarized the mechanisms of ARG resistance, their spread in human and animal populations, and primary input pathways into water bodies, including medical and aquaculture wastewater. Key environmental drivers of ARG evolution and dissemination are analyzed, encompassing selective pressures from antibiotics and heavy metals, horizontal gene transfer mediated by mobile genetic elements, and physicochemical factors such as dissolved oxygen, pH, and nutrient levels. Finally, current research gaps are highlighted, and future directions are proposed for monitoring and controlling ARG transmission, and assessing associated health risks under the One Health framework. This review provides a scientific basis for understanding the ARG crisis in aquatic environments and guiding integrated management strategies.
摘要
抗生素抗性基因(ARGs)作为一种新兴环境污染物,因其在水环境中的持久性、迁移性和传播性而对公众健康构成严重威胁。本文系统综述了ARGs的抗性机制及其在人类和动物群体中的传播,分析了医疗废水、水产养殖废水等主要输入途径。深入探讨了驱动ARGs演化与传播的关键环境因素,包括抗生素和重金属的选择压力、可移动遗传元件介导的水平基因转移,以及溶解氧、pH和营养水平等理化因子。最后,指出了当前研究存在的不足,并在"One Health"框架下提出了未来监测与控制ARGs传播、评估健康风险的研究方向。本综述为理解水环境中ARGs的污染危机及制定综合防控策略提供了科学依据。
Key words: antibiotic resistance genes / transmission mechanisms / aquatic environment drivers / One Health
关键字 : 抗生素抗性基因 / 传播机制 / 水生环境驱动因素 / One Health
Cite this article:SHENG Xuran, SONG Dan, PU Wenmiao, et al. Antibiotic Resistance Genes in Aquatic Environments: Sources, Transmission, and Influencing Factors from a One Health Perspective[J]. Wuhan Univ J of Nat Sci, 2026, 31(2): 146-156.
Biography: SHENG Xuran, female, Master candidate, research direction: distribution and dissemination of antibiotic resistance genes. Email: This email address is being protected from spambots. You need JavaScript enabled to view it.
Foundation item: Supported by the National Natural Science Foundation of China (31802298), and Heilongjiang Postdoctoral Fund of Heilongjiang Province (LBH-Z23303)
© Wuhan University 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
0 Introduction
The discovery and widespread use of antibiotics represent a landmark achievement in modern medicine, revolutionizing the treatment of infectious diseases. However, decades of extensive and often indiscriminate antibiotic application have exerted strong selective pressure on microorganisms, facilitating the rapid evolution and dissemination of antibiotic-resistant bacteria. This adaptive process, while a natural response to antimicrobial exposure, is undermining the foundations of modern healthcare and transforming previously curable infections into potentially lethal threats. Studies suggest that by 2050, antimicrobial resistance (AMR) could become the leading cause of death globally, surpassing mortality from malaria, HIV, and tuberculosis[1]. As such, AMR is now recognized as a major public health crisis worldwide.
China plays a pivotal role in this global challenge. As both a leading producer and consumer of antibiotics, and the world’s largest supplier of livestock and aquaculture products, China’s antibiotic use patterns are closely linked to its agricultural and industrial structures. According to a recent study, the annual growth rate of antibiotic consumption in China has reached 7%, placing it among the highest globally and surpassing that of 23 other upper-middle-income countries[2]. This increasing trend is closely associated with the substantial demand for antibiotics driven by China’s animal husbandry sector. Notably, more than half of the antibiotics produced in China are reportedly used in animal husbandry[3]. Although the prophylactic and therapeutic use of antibiotics in livestock and aquaculture enhances animal health and productivity, it also promotes the accumulation of antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs) within farming environments, forming the initial link in resistance dissemination from agricultural sources to the surrounding environments[4].
Antibiotics and ARGs originating from medical, agricultural, and domestic activities ultimately enter the environment through diverse pathways such as wastewater discharge, manure application, and runoff. Among these pollutants, ARGs have emerged as novel environmental contaminants of global concern due to their ability to be horizontally transferred among microbial populations[5]. ARGs have been extensively detected in multiple environmental matrices, including water[6], soil[7], and even air[8], with aquatic environments serving as reservoirs and transmission hubs. Owing to their fluidity and capacity to integrate pollutants, aquatic systems provide ideal conditions for the accumulation, transformation, and spread of ARGs. A comparative study has shown that water bodies harbor a higher diversity of unique ARGs (32 types) than either sediments (11 types) or soils (5 types)[9]. The persistence and cross-species transfer of ARGs in aquatic environments substantially accelerate the dissemination of resistance, establishing a direct link between environmental reservoirs and clinical infections along the "environment-animal-human" continuum. Of particular concern, the Organization for Economic Co-operation and Development predicts that global antibiotic resistance will double by 2035 compared with the level in 2005[10].
In this context, the One Health framework that integrates human, animal, and environmental health offers a comprehensive and systematic approach to understanding and addressing the AMR crisis. To develop effective control and prevention strategies, it is imperative to systematically elucidate the sources, dissemination routes, and driving mechanisms of ARGs, particularly within aquatic environments. As a pivotal hub, how the aquatic environment receives, mixes, and modulates the flow of ARGs among the microbiomes of humans, animals, and the environment is a key question. To address this question, this review aims to: (1) summarize the major input pathways of ARGs from anthropogenic and animal origins into aquatic environments; (2) explore the synergistic interactions among multiple driving factors, including antibiotics, heavy metals, mobile genetic elements (MGEs), and environmental variables; and (3) provide a scientific foundation for risk assessment and targeted management strategies under the One Health perspective.
1 Mechanisms of Antibiotic Resistance
1.1 Intrinsic Resistance in Natural Populations
Intrinsic resistance refers to the inherent presence of ARG prototypes, quasi-resistance genes, or unexpressed resistance determinants within bacterial genomes. This mechanism constitutes an important natural source of ARGs in the environment[11]. In natural ecosystems, microorganisms gain ecological advantages by synthesizing natural antibiotics while simultaneously evolving intrinsic resistance mechanisms to protect themselves from their own products. Under continuous natural selection process, strains harboring resistance genes acquire evolutionary advantages, whereas susceptible strains are gradually eliminated. Over long evolutionary timescales, such selective forces have not only facilitated the formation and maintenance of ARGs but also contributed to the complexity of modern clinical antimicrobial resistance. A representative example is the emergence and dissemination of resistance in Gram-negative bacteria, which significantly increases infection morbidity and imposes substantial healthcare burdens[12].
Metagenomic investigations have provided robust evidence for the ancient and natural origin of ARGs. Diverse resistance genes have been identified in pristine, human-unaffected environments such as Arctic and Antarctic soils[13], deep permafrost layers[14], and glacial ice[15], demonstrating that antibiotic resistance long predates anthropogenic antibiotic use. Notably, β-lactam, tetracycline, and glycopeptide resistance genes have been detected in approximately 30 000-year-old Beringian permafrost sediments, along with the widespread occurrence of ARGs in isolated cave microbial communities[14, 16]. These findings collectively confirm that ARGs are ancient genetic elements shaped by evolutionary adaptation. They highlight that ARGs are intrinsic products of microbial evolution and reflect the long-standing natural selection pressure imposed by antibiotics within the microbial biosphere. While intrinsic resistance provides the foundational gene pool, the evolutionary success and contemporary crisis of AMR are largely driven by the proliferation and dissemination of these ARGs.
1.2 Proliferation and Dissemination of ARGs
The proliferation and dissemination of ARGs in the environment are primarily driven by two mechanisms: horizontal gene transfer (HGT) and vertical gene transfer (VGT) (Fig. 1). These processes play complementary roles, with HGT enabling rapid cross-species transmission and VGT ensuring the long-term maintenance of resistance within bacterial populations.
 |
Fig. 1 Mechanism of vertical (a) and horizontal (b) transfer of ARGs |
Through HGT, ARGs are mobilized to new bacterial hosts via MGEs, which serve as key vehicles for genetic exchange within and between microbial communities[17]. HGT occurs mainly through conjugation, transduction, and transformation. In conjugation, donor and recipient cells establish direct contact through pili, forming a transfer channel that mediates the exchange of plasmid or chromosomal DNA carrying ARGs[18]. Conjugative elements often harbor multiple resistance genes and represent the predominant route of ARG dissemination. For instance, horizontal transfer of the conjugative plasmid pKJK5, carrying the trimethoprim resistance gene (dfrA1) and tetA, has been observed within biofilms formed by diverse bacterial species on plastic surfaces in aquatic environments[19]. Transduction refers to bacteriophage-mediated transfer of ARGs from donor to recipient bacteria, typically conferring resistance to a specific antibiotic class[20]. Phages can carry beneficial genes that enhance host fitness, thereby promoting their own persistence. ARGs have been identified in phage fractions from hospital and wastewater treatment plant effluents, as well as from animal and human fecal samples, by qPCR detection methods[21]. These findings suggest that bacteriophages act as important reservoirs and vectors of ARGs in natural and engineered environments. Transformation involves the uptake of extracellular DNA fragments from lysed resistant cells by competent bacterial hosts, followed by genomic integration and gene expression that confer antibiotic resistance[22]. Genomic analysis of soil Listeria has shown that the sensu stricto lineage possesses more intact competence genes than the sensu lato group. Key genes (e.g., comK, comEC) are over 90% functional in sensu stricto, while others (e.g., comGD, comGF) are absent in sensu lato. Genomes with recombinant ARGs carry significantly more functional competence genes (an average of 11)[23] , indicating natural transformation is a predominant pathway for ARG transfer in these bacteria. Taken together, these HGT processes facilitate the rapid spread of resistance traits across taxonomic and ecological boundaries, greatly complicating the management of infectious diseases.
In contrast, VGT refers to the inheritance of ARGs from parent to offspring cells during bacterial replication, leading to the stable amplification of resistance genes within clonal lineages[24]. In structured microbial communities, such as biofilms, sediments, or microplastic-associated assemblages, VGT contributes to the persistence and ecological stability of ARGs. For instance, riverine microplastics can act as colonization surfaces for ARB, thereby enhancing the proliferation of ARGs through vertical transmission within these microhabitats[25]. Although research on ARG dissemination has primarily focused on HGT, recent advances highlight the complementary role of VGT under specific environmental conditions. Microfluidic analysis of activated sludge communities exposed to trimethoprim, tetracycline, and amoxicillin revealed that while antibiotic action on donors and recipients modulates HGT efficiency, VGT plays a crucial role in maintaining transconjugant populations and sustaining ARG persistence[26]. Additionally, low-density polyethylene facilitates the vertical transfer of ARGs by increasing bacterial cell proliferation and enhancing the expression of relevant functional genes[27]. Thus, despite its slower interspecies spread, VGT serves as an essential mechanism for conserving resistance traits and sustaining the environmental resistome.
Antibiotic resistance dissemination involves two synergistic mechanisms: HGT acts as an innovator, randomly spreading resistance genes across bacterial species via MGEs. VGT then serves as an amplifier, enabling the clonal expansion of bacteria that have acquired these genes. This combination of HGT-mediated diversification and VGT-driven amplification powerfully accelerates the spread of resistance.
2 Prevalence and Dissemination of ARGs in Human and Animal Populations
To date, thousands of antibiotics, including natural, semi-synthetic, and fully synthetic compounds, have been discovered. Their antimicrobial spectra have expanded beyond antibacterial effects to include antiviral, antitumor, and antiparasitic activities[28]. The extensive use of antibiotics in human medicine and animal husbandry has shaped distinct resistomes, which are interconnected through multiple transmission pathways across ecological boundaries.
In clinical settings, β-lactam antibiotics represent one of the most extensively used antimicrobial classes owing to their broad activity spectrum and clinical effectiveness, particularly against bloodstream infections and other severe conditions[29]. Escherichia coli (E. coli), a common commensal bacterium in the intestinal tracts of humans and animals, exhibits a strong capacity to carry and disseminate ARGs in both clinical and environmental contexts. Studies have confirmed that E. coli isolates from hospital effluents, drinking water, and food chains harbor a variety of β-lactamase genes (including blaCTX-M-9, blaCTX-M-15, and blaTEM) as well as plasmid-mediated quinolone resistance genes such as qnrS[30-31]. These resistant bacteria circulate through aquatic and food systems, facilitating ARG dissemination across ecological compartments. Once colonizing humans, they can cause diarrhea, urinary tract infections, bloodstream infections, and nosocomial bacteremia, posing severe challenges to clinical anti-infective treatment[30-31]. As a key vector for β-lactam ARGs, the continuous circulation of E. coli between the environmental and clinical systems reinforces resistance transmission networks and underscores potential pathways for resistant bacteria to move from the environment to humans, thereby aggravating the public health burden of antibiotic resistance. The resistance issue associated with Staphylococcus aureus (S. aureus) is equally concerning. This pathogen demonstrates remarkable adaptability and high levels of resistance in both hospital and community environments[32]. Studies have frequently detected resistance genes such as mecA, tetK, and blaZ in S. aureus isolates derived from food sources[33-34], confirming its broad resistance to β-lactams and tetracyclines. Particularly noteworthy is Methicillin-resistant S. aureus (MRSA), a classic hospital-acquired pathogen whose resistance to β-lactam antibiotics greatly complicates treatment and increases patient mortality risk[34].
Animal farming also plays a pivotal role in ARG transmission. The misuse of antibiotics for the prevention and treatment of bovine mastitis caused by Klebsiella pneumoniae has led to the detection of multiple ARGs in milk, heightening the risk of resistance dissemination[35]. Research on Klebsiella oxytoca in swine production systems has shown high detection rates for blaTEM, blaCTX-M, and blaSHV β-lactamase genes, suggesting their dominance among pig-derived strains and reflecting the extensive presence and transmission potential of ARGs in livestock production chains[36].
Aquaculture similarly constitutes a major pathway for ARG propagation. Enterococcus, a common gut bacterium in aquatic animals, has been found in Nile tilapia to carry vancomycin resistance genes (vanA and vanC), exhibiting multidrug resistance and pathogenicity to fish. These findings suggest that aquaculture may serve as an additional route for ARG transmission to humans[37].
Environmental media are also crucial in determining the fate and redistribution of ARGs, with aquatic systems in particularly serving as hotspots for resistance gene exchange among bacteria. Salmonella, a typical foodborne pathogen, has been widely detected in water environments. For instance, monitoring of Jordan’s two major irrigation sources, the King Abdullah Canal and Wadi Al-Wala, revealed that 32.2% of samples were contaminated with Salmonella, with some strains showing resistance to tetracycline and other antibiotics[38]. This indicates the potential role of agricultural irrigation water in transferring resistant Salmonella to crops and, ultimately, to humans[38-39]. Pseudomonas aeruginosa, a ubiquitous species in natural aquatic environments such as lakes and rivers, displays multidrug resistance to aminoglycosides, β-lactams, and quinolones. Environmental isolates have been found to harbor multiple ARGs, including sul1, qnrS, blaVIM, blaTEM, blaCTX, blaAIM-1, tetA, and ampC[40].
In summary, the bidirectional transmission of ARGs between human and animal veterinary sectors constitutes a critical global health concern. Infections caused by multidrug-resistant bacteria not only heighten disease burdens but also create "untreatable" therapeutic challenges, emphasizing the urgency and complexity of addressing the current antibiotic resistance crisis.
3 Anthropogenic Sources of ARGs in Aquatic Environments
The origins of ARGs in aquatic environments are complex, arising from interactions between anthropogenic activities and natural microbial ecosystems. Key contributors include healthcare systems, aquaculture, and agricultural practices, which serve as primary sources of exogenous ARG input (Fig. 2).
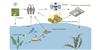 |
Fig. 2 Main anthropogenic source and spread of ARGs in aquatic environment |
3.1 Medical Wastewater Discharge and Clinical Antibiotic Residues
Healthcare systems constitute a major source of ARG generation and dissemination. Bacteria isolated from hospital environments frequently exhibit antibiotic resistance. Antibiotics are extensively used in medical institutions, yet approximately 80%-90% of administered compounds are excreted by patients as parent drugs or active metabolites into wastewater via feces and urine[41]. Monitoring studies from six tertiary hospitals in northern India identified 1 573 microbial species and 885 ARG subtypes in medical wastewater, with Proteobacteria carrying 85% of detected ARGs and serving as dominant ARG reservoirs[42].
High microbial density in hospital wastewater further facilitates contact among resistant bacterial species, substantially increasing the risk of ARG transmission through HGT. Metagenomic analyses have revealed that medical wastewater contains diverse ARGs, including aac(3), blaOXA, and blaTEM, with abundance and diversity exceeding those observed in other wastewater systems[42]. A large-scale study revealed that although the ARG load in the discharged effluents of hospital wastewater systems (0.58-5.91 copies/cell; 175-448 distinct subtypes) was lower than that in the influents (0.96-7.94 copies/cell; 205-539 subtypes), they still carried a substantial pollutant burden[43]. These effluents, containing antibiotic residues, metabolites, and resistant bacteria, enter surrounding environments through discharge systems, representing important pathways for ARG contamination in aquatic environments.
3.2 Continuous Input from Aquaculture and Veterinary Medicine
Aquaculture facilities and veterinary sites represent additional hotspots for ARG enrichment and spread. High-density aquaculture systems are prone to infectious outbreaks, leading to the widespread use of antibiotics as prophylactics, feed additives, and growth promoters to improve production efficiency[44]. Unmetabolized antibiotics from feed ultimately accumulate in water and sediments[45], driving the evolution of multidrug-resistant bacteria[46]. ARB and ARGs can be transmitted through the food chain, moving from animal hosts to other environmental media or higher trophic levels, thereby expanding the dissemination range[47-48]. Consumption of ARG-contaminated animal products subsequently poses direct risks to human health and increases public healthcare burdens.
Intensive livestock operations in swine, poultry, and cattle production have also substantially increased antibiotic demand. A considerable fraction of these antibiotics persists in manure, which is applied to agricultural land as fertilizer. Subsequent runoff from manure-amended fields during rainfall events transports antibiotics and ARGs to aquatic systems, contributing to environmental resistance genes dissemination[49]. This non-point source pollution further contributes to the widespread contamination of aquatic environments, transforming them into both reservoirs and dissemination vectors for antibiotic resistance. Consequently, water bodies serve as a crucial nexus connecting agricultural antibiotic use to broader environmental and public health exposure.
3.3 Agricultural Runoff and Pesticide-Mediated Drivers
Agricultural practices constitute another significant source of ARGs in water bodies. Rising global food demands have led to increased application of pesticides, including herbicides, fungicides, and insecticides. A study demonstrates that pesticides can promote ARG enrichment through cross-resistance mechanisms between pesticide-resistant bacteria and antibiotic-resistant bacteria, as well as via plasmid-mediated gene transfer[50]. The transport of ARGs from farmland to aquatic ecosystems exhibits distinct seasonal patterns: during the rainy season, heavy rainfall generates substantial surface runoff that carries higher loads of ARGs and pesticides into water bodies, while, in dry seasons, limited water flow results in lower but more consistent baseline contamination[51]. Certain pesticides, particularly insecticides, can modulate bacterial stress responses and influence resistance acquisition, further enhancing the development of antibiotic resistance in aquatic microbiomes[20]. In summary, agricultural non-point source pollution plays a significant role in ARG input to freshwater ecosystems.
4 Factors Influencing ARGs in Aquatic Environments
Aquatic environments serve as critical reservoirs for ARGs, hosting diverse pollutants including antibiotics, heavy metals, and microorganisms in untreated waters (Fig. 3). These factors collectively shape the emergence, persistence, and dissemination of both ARGs and ARB.
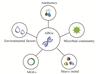 |
Fig. 3 Main influence factors of ARGs in aquatic environment |
4.1 Direct Selective Pressure from Antibiotics
Continuous introduction of antibiotics into aquatic systems generates "pseudo-persistent" pollution, directly inducing the emergence and spread of ARGs and ARB[52-54]. Selective pressure exerted by antibiotics is a primary driver of bacterial resistance development[55]. However, the relationship between ARG abundance and antibiotic concentrations is not uniformly consistent. Positive correlations have been observed, such as between tetracycline antibiotics and tetA, tetB, and tetC in Honghu Lake[56], or between total antibiotic concentrations and sul1, sul2, and total ARGs levels in Poyang Lake during the rainy season[57]. Conversely, no significant correlations were detected in water bodies such as Fuxian Lake and Dongting Lake[58-59]. This inconsistency reflects the complex interplay of multiple factors beyond antibiotic presence alone, including variations in microbial community composition, the differential mobilization of ARGs via MGEs, and seasonal environmental changes that collectively modulate ARG dynamics and transfer efficiency.
4.2 MGEs
HGT mediated by MGEs is a key mechanism for ARG proliferation. MGEs, including plasmids, integrons, transposons, insertion sequences, genomic islands, and bacteriophages, serve as vectors for ARG transfer among bacteria, enhancing dissemination and adaptation under environmental stress[53, 60-62]. Class 1 integrons capture and express ARGs and are strongly associated with sulfonamide resistance genes (sul1 and sul2) in surface waters and sediments[54, 63]. Plasmids facilitate cross-species ARG transmission, as evidenced by the dissemination of blaOXA-48-like genes in carbapenem-resistant Klebsiella pneumoniae across Europe[64]. Detection rates of MGEs, such as tnpA, show positive correlations with specific ARGs (e.g., bacA) in the Yellow River[65], suggesting that MGEs can serve as indicators of ARG potential and are crucial targets for resistance management[66].
4.3 Heavy Metal Co-Selection Mechanisms
Heavy metals, persistent and bioaccumulative pollutants originating from mining, agriculture, and aquaculture, exert continuous selective pressure on microbial communities, promoting ARG development[67-68]. Mechanisms include cross-resistance, co-resistance, and shared regulatory pathways, such as efflux pumps or cell wall modification that confer simultaneous resistance to metals and antibiotics[67, 69-70]. Low-level metal exposure can enhance integron activity (e.g., intI1), facilitating HGT and dissemination, whereas excessive concentrations may inhibit ARGs proliferation[63, 71]. Co-occurrence of ARGs and metal resistance genes (MRGs) increases pathogenic potential, as seen with copper and zinc stress enhances the abundance of MRGs (czcD, pocA, copB) and ARGs (sul2, ermB, tetA)[72-73].
4.4 Environmental Factors
Environmental parameters, including dissolved oxygen (DO), water temperature (WT), pH, salinity, total nitrogen (TN), and total phosphorus (TP), indirectly or directly influence ARG distribution and transmission. Hypoxic conditions promote HGT[74], while elevated temperatures increase bacterial growth and ARG proliferation, enhancing abundances of tetM, mecA, bacA, vatE, and tetW[75]. Optimal pH conditions can favor ARG expression in specific bacterial taxa[76]. Nutrient enrichment, particularly in eutrophic systems, stimulates microbial growth and promotes ARG transmission, as observed in Chaohu Lake and Honghu Lake[56, 77-79].
4.5 Microbial Community Structure
Microbial communities act as natural hosts and vectors for ARGs, with community composition strongly influencing ARG persistence and flux in aquatic environments[52]. Dominant phyla such as Proteobacteria, Bacteroidetes, and Actinobacteria are positively correlated with various ARGs[54, 79-80]. In Wanfeng Lake, Proteobacteria were associated with sul1 and tetA, while Actinobacteria correlated with tetA, tetB, tetW, and qnrB[81]. Bacteroidetes, common in eutrophic waters, show positive correlations with multiple ARGs (cmlA, floR, qnrD, qnrS, sul1, sul2, tetG, tetX, tetW) [82]. Pathogenic genera, including Acinetobacter, Burkholderia, and Pseudomonas, serve as ARG reservoirs, highlighting their potential risks to human health and the importance of understanding microbial community and ARG interactions for early warning and mitigation strategies[79].
5 Conclusions and Perspectives
This review delineates the complex "source-transmission-regulation" network governing ARGs in aquatic environments, emphasizing the critical role of anthropogenic inputs combined with genetic and environmental drivers.
Tackling aquatic ARG pollution necessitates innovative strategies beyond conventional approaches. Key research priorities should include:
1) Predictive framework development: Integrate multi-omics data with machine learning to forecast ARG dissemination risks and identify potential intervention points.
2) High-risk pathogen identification: Apply advanced cultivation techniques and single-cell genomics to resolve clinically significant ARG-host-vector combinations within complex communities.
3) Horizontal gene transfer inhibition: Developing targeted strategies to disrupt conjugation and transformation processes, potentially through specific molecular inhibitors or phage-based interventions.
4) Engineered barrier design: Creating treatment systems and constructed ecosystems that not only remove contaminants but actively suppress ARG mobility through optimized microbial selection and redox manipulation.
Advancing these fronts will enable a crucial transition from passive monitoring to active intervention, ultimately strengthening our capacity to safeguard water resources and public health against the escalating threat of AMR.
References
- Painter C, Limmathurotsakul D, Roberts T, et al. Sustainable antimicrobial resistance surveillance: Time for a global funding mechanism[J]. The Lancet Infectious Diseases, 2025, 25(2): e99-e103. [Google Scholar]
- Wang L G, Chen H, Zhang Y Y, et al. Global antibiotic consumption and regional antimicrobial resistance, 2010-21: An analysis of pharmaceutical sales and antimicrobial resistance surveillance data[J]. The Lancet Global Health, 2025, 13(11): e1880-e1891. [Google Scholar]
- Shao Y T, Wang Y P, Yuan Y W, et al. A systematic review on antibiotics misuse in livestock and aquaculture and regulation implications in China[J]. Science of the Total Environment, 2021, 798: 149205. [Google Scholar]
- Zhi S L, Shen S Z, Zhou J, et al. Systematic analysis of occurrence, density and ecological risks of 45 veterinary antibiotics: Focused on family livestock farms in Erhai Lake basin, Yunnan, China[J]. Environmental Pollution, 2020, 267: 115539. [Google Scholar]
- Pruden A, Pei R T, Storteboom H, et al. Antibiotic resistance genes as emerging contaminants: Studies in northern Colorado[J]. Environmental Science & Technology, 2006, 40(23): 7445-7450. [Google Scholar]
- Chen Y H, Su J Q, Zhang J Y, et al. High-throughput profiling of antibiotic resistance gene dynamic in a drinking water river-reservoir system[J]. Water Research, 2019, 149: 179-189. [Google Scholar]
- Guo M Y, Wu Y F, Huang H Y, et al. Revealing the critical role of rare bacterial communities in shaping antibiotic resistance genes in saline soils through metagenomic analysis[J]. Journal of Hazardous Materials, 2025, 491: 137848. [Google Scholar]
- Wang Y, Han Y P, Li L, et al. Airborne human-associated ARGs in municipal wastewater treatment plants[J]. Journal of Hazardous Materials, 2025, 498: 139766. [Google Scholar]
- Chen Y H, Li P, Huang Y S, et al. Environmental media exert a bottleneck in driving the dynamics of antibiotic resistance genes in modern aquatic environment[J]. Water Research, 2019, 162: 127-138. [Google Scholar]
- Leung E, Weil D E, Raviglione M, et al. The WHO policy package to combat antimicrobial resistance[J]. Bulletin of the World Health Organization, 2011, 89(5): 390-392. [Google Scholar]
- Davies J, Davies D. Origins and evolution of antibiotic resistance[J]. Microbiology and Molecular Biology Reviews, 2010, 74(3): 417-433. [Google Scholar]
- Das S, Bush K. Success and challenges associated with large-scale collaborative surveillance for carbapenemase genes in gram-negative bacteria[J]. Antimicrobial Agents and Chemotherapy, 2022, 66(2): e02299-e02221. [Google Scholar]
- Van Goethem M W, Pierneef R, Bezuidt O K I, et al. A reservoir of 'historical' antibiotic resistance genes in remote pristine Antarctic soils[J]. Microbiome, 2018, 6(1): 40. [Google Scholar]
- D'Costa V M, King C E, Kalan L, et al. Antibiotic resistance is ancient[J]. Nature, 2011, 477(7365): 457-461. [Google Scholar]
- Sajjad W, Ilahi N, Haq A, et al. Bacteria populating freshly appeared supraglacial lake possess metals and antibiotic-resistant genes[J]. Environmental Research, 2024, 247: 118288. [Google Scholar]
- Bhullar K, Waglechner N, Pawlowski A, et al. Antibiotic resistance is prevalent in an isolated cave microbiome[J]. PLoS One, 2012, 7(4): e34953. [Google Scholar]
- Shao S C, Hu Y Y, Cheng J H, et al. Research progress on distribution, migration, transformation of antibiotics and antibiotic resistance genes (ARGs) in aquatic environment[J]. Critical Reviews in Biotechnology, 2018, 38(8): 1195-1208. [Google Scholar]
- Sunde M, Norström M. The prevalence of, associations between and conjugal transfer of antibiotic resistance genes in Escherichia coli isolated from Norwegian meat and meat products[J]. The Journal of Antimicrobial Chemotherapy, 2006, 58(4): 741-747. [Google Scholar]
- Arias-Andres M, Klümper U, Rojas-Jimenez K, et al. Microplastic pollution increases gene exchange in aquatic ecosystems[J]. Environmental Pollution, 2018, 237: 253-261. [Google Scholar]
- Balcazar J L. Bacteriophages as vehicles for antibiotic resistance genes in the environment[J]. PLoS Pathogens, 2014, 10(7): e1004219. [Google Scholar]
- Marti E, Variatza E, Balcázar J L. Bacteriophages as a reservoir of extended-spectrum β-lactamase and fluoroquinolone resistance genes in the environment[J]. Clinical Microbiology and Infection, 2014, 20(7): O456-O459. [Google Scholar]
- Krzeminski P, Feys E, Anglès d'Auriac M, et al. Combined membrane filtration and 265 nm UV irradiation for effective removal of cell free antibiotic resistance genes from feed water and concentrate[J]. Journal of Membrane Science, 2020, 598: 117676. [Google Scholar]
- Goh Y X, Anupoju S M B, Nguyen A, et al. Evidence of horizontal gene transfer and environmental selection impacting antibiotic resistance evolution in soil-dwelling Listeria[J]. Nature Communications, 2024, 15: 10034. [Google Scholar]
- Joakim Larsson D G, Flach C F. Antibiotic resistance in the environment[J]. Nature Reviews Microbiology, 2022, 20(5): 257-269. [Google Scholar]
- Wang S S, Xue N N, Li W F, et al. Selectively enrichment of antibiotics and ARGs by microplastics in river, estuary and marine waters[J]. Science of the Total Environment, 2020, 708: 134594. [Google Scholar]
- Li B, Qiu Y, Song Y Q, et al. Dissecting horizontal and vertical gene transfer of antibiotic resistance plasmid in bacterial community using microfluidics[J]. Environment International, 2019, 131: 105007. [Google Scholar]
- Luo T Y, Dai X H, Wei W, et al. Microplastics enhance the prevalence of antibiotic resistance genes in anaerobic sludge digestion by enriching antibiotic-resistant bacteria in surface biofilm and facilitating the vertical and horizontal gene transfer[J]. Environmental Science & Technology, 2023, 57(39): 14611-14621. [Google Scholar]
- Leisner J J. The diverse search for synthetic, semisynthetic and natural product antibiotics from the 1940s and up to 1960 exemplified by a small pharmaceutical player[J]. Frontiers in Microbiology, 2020, 11: 976. [Google Scholar]
- De Angelis G, Del Giacomo P, Posteraro B, et al. Molecular mechanisms, epidemiology, and clinical importance of β-lactam resistance in Enterobacteriaceae[J]. International Journal of Molecular Sciences, 2020, 21(14): 5090. [Google Scholar]
- Talukdar P K, Rahman M, Rahman M, et al. Antimicrobial resistance, virulence factors and genetic diversity of Escherichia coli isolates from household water supply in Dhaka, Bangladesh[J]. PLoS One, 2013, 8(4): e61090. [Google Scholar]
- Rasheed M U, Thajuddin N, Ahamed P, et al. Antimicrobial drug resistance in strains of Escherichia coli isolated from food sources[J]. Revista Do Instituto de Medicina Tropical de Sao Paulo, 2014, 56(4): 341-346. [Google Scholar]
- Ahmad-Mansour N, Loubet P, Pouget C, et al. Staphylococcus aureus toxins: An update on their pathogenic properties and potential treatments[J]. Toxins, 2021, 13(10): 677. [Google Scholar]
- Jia K, Fang T S, Wang X, et al. Antibiotic resistance patterns of staphylococcus aureus isolates from retail foods in mainland China: A meta-analysis[J]. Foodborne Pathogens and Disease, 2020, 17(5): 296-307. [Google Scholar]
- Mkize N, Zishiri O T, Mukaratirwa S. Genetic characterisation of antimicrobial resistance and virulence genes in Staphylococcus aureus isolated from commercial broiler chickens in the Durban metropolitan area, South Africa[J]. Journal of the South African Veterinary Association, 2017, 88: 1416. [Google Scholar]
- Kang H J, You J Y, Kim S H, et al. Genetic diversity, virulence genes, antimicrobial resistance, and biofilm formation of Klebsiella pneumoniae isolated from bovine mastitis milk in South Korea[J]. Microbiology Spectrum, 2025, 13(11): e01343-25. [Google Scholar]
- Phetburom N, Boueroy P, Chopjitt P, et al. Phenotypic and molecular characterization of β-lactamase and plasmid-mediated quinolone resistance genes in Klebsiella oxytoca isolated from slaughtered pigs in Thailand[J]. Veterinary World, 2022, 15(2): 309-315. [Google Scholar]
- Osman K M, Ali M N, Radwan I, et al. Dispersion of the vancomycin resistance genes vanA and vanC of enterococcus isolated from Nile tilapia on retail sale: A public health hazard[J]. Frontiers in Microbiology, 2016, 7: 1354. [Google Scholar]
- Burjaq S Z, Abu-Romman S M. Prevalence and antimicrobial resistance of Salmonella spp. from irrigation water in two major sources in Jordan[J]. Current Microbiology, 2020, 77(11): 3760-3766. [Google Scholar]
- Bell R L, Jarvis K G, Ottesen A R, et al. Recent and emerging innovations in Salmonella detection: A food and environmental perspective[J]. Microbial Biotechnology, 2016, 9(3): 279-292. [Google Scholar]
- Reem A, Almansoob S, Senan A M, et al. Pseudomonas aeruginosa and related antibiotic resistance genes as indicators for wastewater treatment[J]. Heliyon, 2024, 10(9): e29798. [Google Scholar]
- Fernanda P A, Liu S, Yuan T M, et al. Diversity and abundance of antibiotic resistance genes and their relationship with nutrients and land use of the inflow rivers of Taihu Lake[J]. Frontiers in Microbiology, 2022, 13: 1009297. [Google Scholar]
- Talat A, Blake K S, Dantas G, et al. Metagenomic insight into microbiome and antibiotic resistance genes of high clinical concern in urban and rural hospital wastewater of northern India origin: A major reservoir of antimicrobial resistance[J]. Microbiology Spectrum, 2023, 11(2): e04102-22. [Google Scholar]
- Shuai X Y, Zhou Z C, Zhu L, et al. Ranking the risk of antibiotic resistance genes by metagenomic and multifactorial analysis in hospital wastewater systems[J]. Journal of Hazardous Materials, 2024, 468: 133790. [Google Scholar]
- Carvalho Pereira J, Lemoine A, Neubauer P, et al. Perspectives for improving circular economy in brackish shrimp aquaculture[J]. Aquaculture Research, 2022, 53(4): 1169-1180. [Google Scholar]
- Xu M, Huang X H, Shen X X, et al. Metagenomic insights into the spatiotemporal responses of antibiotic resistance genes and microbial communities in aquaculture sediments[J]. Chemosphere, 2022, 307: 135596. [Google Scholar]
- Suyamud B, Lohwacharin J, Yang Y Y, et al. Antibiotic resistant bacteria and genes in shrimp aquaculture water: Identification and removal by ferrate(Ⅵ)[J]. Journal of Hazardous Materials, 2021, 420: 126572. [Google Scholar]
- Klous G, Huss A, Heederik D J J, et al. Human-livestock contacts and their relationship to transmission of zoonotic pathogens, a systematic review of literature[J]. One Health, 2016, 2: 65-76. [Google Scholar]
- Beyene T. Veterinary drug residues in food-animal products: Its risk factors and potential effects on public health[J]. Journal of Veterinary Science & Technology, 2015, 7(1): 285. [Google Scholar]
- Gu Y R, Shen S Z, Han B J, et al. Family livestock waste: An ignored pollutant resource of antibiotic resistance genes[J]. Ecotoxicology and Environmental Safety, 2020, 197: 110567. [Google Scholar]
- Shi L M, Zhang J Y, Lu T D, et al. Metagenomics revealed the mobility and hosts of antibiotic resistance genes in typical pesticide wastewater treatment plants[J]. Science of the Total Environment, 2022, 817: 153033. [Google Scholar]
- Yang H, Fu H W, Zhao Y T, et al. Antibiotic resistance genes in multi-matrices of Chaohu Lake: Spatiotemporal variation and correlation with pesticides and PPCPs[J]. Journal of Hazardous Materials, 2025, 497: 139604. [Google Scholar]
- Zhang X X, Zhang T, Fang H H P. Antibiotic resistance genes in water environment[J]. Applied Microbiology and Biotechnology, 2009, 82(3): 397-414. [Google Scholar]
- Zhao R X, Yu K, Zhang J Y, et al. Deciphering the mobility and bacterial hosts of antibiotic resistance genes under antibiotic selection pressure by metagenomic assembly and binning approaches[J]. Water Research, 2020, 186: 116318. [Google Scholar]
- Wang Y Q, Lu S Y, Liu X H, et al. Profiles of antibiotic resistance genes in an inland salt-lake Ebinur Lake, Xinjiang, China: The relationship with antibiotics, environmental factors, and microbial communities[J]. Ecotoxicology and Environmental Safety, 2021, 221: 112427. [Google Scholar]
- Ben W W, Wang J, Cao R K, et al. Distribution of antibiotic resistance in the effluents of ten municipal wastewater treatment plants in China and the effect of treatment processes[J]. Chemosphere, 2017, 172: 392-398. [Google Scholar]
- Wang Z, Han M Z, Li E H, et al. Distribution of antibiotic resistance genes in an agriculturally disturbed lake in China: Their links with microbial communities, antibiotics, and water quality[J]. Journal of Hazardous Materials, 2020, 393: 122426. [Google Scholar]
- Liang X M, Guan F L, Chen B W, et al. Spatial and seasonal variations of antibiotic resistance genes and antibiotics in the surface waters of Poyang Lake in China[J]. Ecotoxicology and Environmental Safety, 2020, 196: 110543. [Google Scholar]
- Yang Y Y, Cao X H, Lin H, et al. Antibiotics and antibiotic resistance genes in sediment of Honghu Lake and east Dongting Lake, China[J]. Microbial Ecology, 2016, 72(4): 791-801. [Google Scholar]
- Zhao B, Xu J M, Zhang G D, et al. Occurrence of antibiotics and antibiotic resistance genes in the Fuxian Lake and antibiotic source analysis based on principal component analysis-multiple linear regression model[J]. Chemosphere, 2021, 262: 127741. [Google Scholar]
- Samreen, Ahmad I, Malak H A, et al. Environmental antimicrobial resistance and its drivers: A potential threat to public health[J]. Journal of Global Antimicrobial Resistance, 2021, 27: 101-111. [Google Scholar]
- Tokuda M, Shintani M. Microbial evolution through horizontal gene transfer by mobile genetic elements[J]. Microbial Biotechnology, 2024, 17(1): e14408. [Google Scholar]
- Ghaly T M, Gillings M R. New perspectives on mobile genetic elements: A paradigm shift for managing the antibiotic resistance crisis[J]. Philosophical Transactions of the Royal Society B: Biological Sciences, 2022, 377(1842): 20200462. [Google Scholar]
- Gupta S, Graham D W, Sreekrishnan T R, et al. Effects of heavy metals pollution on the co-selection of metal and antibiotic resistance in urban rivers in UK and India[J]. Environmental Pollution, 2022, 306: 119326. [Google Scholar]
- Cross B J, Partridge S R, Sheppard A E. Impacts of mobile genetic elements on antimicrobial resistance genes in gram-negative pathogens: Current insights and genomic approaches[J]. Microbiological Research, 2026, 302: 128340. [Google Scholar]
- Wang B, Song L, Li W J, et al. Distribution and migration of antibiotic resistance genes, as well as their correlation with microbial communities in swine farm and its surrounding environments[J]. Environmental Pollution, 2023, 316: 120618. [Google Scholar]
- Yan M T, Xu C, Huang Y M, et al. Tetracyclines, sulfonamides and quinolones and their corresponding resistance genes in the Three Gorges Reservoir, China[J]. Science of the Total Environment, 2018, 631: 840-848. [Google Scholar]
- Imran M, Das K R, Naik M M. Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: An emerging health threat[J]. Chemosphere, 2019, 215: 846-857. [Google Scholar]
- Stepanauskas R, Glenn T C, Jagoe C H, et al. Elevated microbial tolerance to metals and antibiotics in metal-contaminated industrial environments[J]. Environmental Science & Technology, 2005, 39(10): 3671-3678. [Google Scholar]
- Baker-Austin C, Wright M S, Stepanauskas R, et al. Co-selection of antibiotic and metal resistance[J]. Trends in Microbiology, 2006, 14(4): 176-182. [Google Scholar]
- Zhang N, Juneau P, Huang R L, et al. Coexistence between antibiotic resistance genes and metal resistance genes in manure-fertilized soils[J]. Geoderma, 2021, 382: 114760. [Google Scholar]
- Li H, Xu H, Song H L, et al. Antibiotic resistance genes, bacterial communities, and functions in constructed wetland-microbial fuel cells: Responses to the co-stresses of antibiotics and zinc[J]. Environmental Pollution, 2020, 265: 115084. [Google Scholar]
- Nnaji N D, Anyanwu C U, Miri T, et al. Mechanisms of heavy metal tolerance in bacteria: A review[J]. Sustainability, 2024, 16(24): 11124. [Google Scholar]
- Wei H D, Wu X, Chen A X, et al. Reserve and proliferation characteristics of antibiotic resistance genes and heavy metal resistance genes in the sewage pipe biofilm under the stress of Cu and Zn[J]. Journal of Environmental Chemical Engineering, 2025, 13(2): 116162. [Google Scholar]
- Deng Y Q, Jiang J J, Huang Y B, et al. Hypoxia triggers the proliferation of antibiotic resistance genes in a marine aquaculture system[J]. Science of the Total Environment, 2023, 859: 160305. [Google Scholar]
- Yu Q L, Han Q, Shi S Q, et al. Metagenomics reveals the response of antibiotic resistance genes to elevated temperature in the Yellow River[J]. Science of the Total Environment, 2023, 859(Pt 2): 160324. [Google Scholar]
- Liu S G, Xu Q H, Lou S, et al. Spatiotemporal distributions of sulfonamide and tetracycline resistance genes and microbial communities in the coastal areas of the Yangtze River Estuary[J]. Ecotoxicology and Environmental Safety, 2023, 259: 115025. [Google Scholar]
- Ohore O E, Qin Z R, Sanganyado E, et al. Ecological impact of antibiotics on bioremediation performance of constructed wetlands: Microbial and plant dynamics, and potential antibiotic resistance genes hotspots[J]. Journal of Hazardous Materials, 2022, 424: 127495. [Google Scholar]
- Luo Y Y, Liu C, Wang Y, et al. Occurrence, distribution and their correlation with different parameters of antibiotics and antibiotic resistance genes in lakes of China: A review[J]. Marine Pollution Bulletin, 2023, 193: 115189. [Google Scholar]
- Han M Z, Zhang L, Zhang N, et al. Antibiotic resistome in a large urban-lake drinking water source in middle China: Dissemination mechanisms and risk assessment[J]. Journal of Hazardous Materials, 2022, 424(Pt D): 127745. [Google Scholar]
- Liu Y Y, Chu K J, Hua Z L, et al. Dynamics of antibiotic resistance genes in the sediments of a water-diversion lake and its human exposure risk behaviour[J]. Science of the Total Environment, 2024, 929: 172563. [Google Scholar]
- Zhang Y T, Li J, Wu T Y, et al. Characteristics of antibiotic resistance genes and microbial community distribution in Wanfeng Lake, upper Pearl River, China[J]. Environmental Science and Pollution Research, 2023, 30(35): 83214-83230. [Google Scholar]
- Li W J, Su H C, Cao Y C, et al. Antibiotic resistance genes and bacterial community dynamics in the seawater environment of Dapeng Cove, South China[J]. Science of the Total Environment, 2020, 723: 138027. [Google Scholar]
All Figures
 |
Fig. 1 Mechanism of vertical (a) and horizontal (b) transfer of ARGs |
| In the text | |
 |
Fig. 2 Main anthropogenic source and spread of ARGs in aquatic environment |
| In the text | |
 |
Fig. 3 Main influence factors of ARGs in aquatic environment |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


